CANDLE-J for denoising 2p data
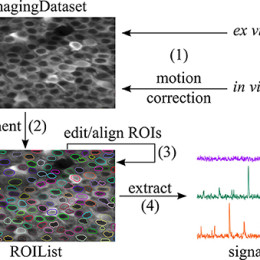
Collaborative Approach for eNhanced Denoising under Low-light Excitation, or CANDLE, is a denoising algorithm specialized for the type of images that are acquired in 2-photon imaging applications. There’s code for both ImageJ and MATLAB available at…